From algorithms to the bench
Text-mining holds the promise of helping researchers to overcome information overload. It is a familiar premise: an avalanche of scientific knowledge is being produced and shared. Teaching machines to “read” might soon be the only manageable way to digest large amounts of information into useful facts.
To extract bits of information, such as biological concepts or relations, from the text, various text-mining tools have been developed in the recent years. Even as the text-mining technology becomes more widespread, an average researcher will rarely be exposed to its benefits. Scholars get to publications via a few familiar routes, which may not utilise text-mining technologies. In addition, different text-mining platforms may focus on different topics or categories: one suited for molecular interactions, another for gene-disease associations.
To address these issues, Europe PMC has established a platform that consolidates text-mined annotations from different sources and makes them available to the wider research community. The annotated concepts and relations are displayed on article pages via SciLite tool, and can be retrieved using RESTful API.
To simplify the process of sharing text-mining results Europe PMC has developed a dedicated Annotation Submission System. It allows expert text-mining providers to publish their annotations in Europe PMC. The system can also accept relevant statements manually curated by dedicated biocuration groups.
The submission process is straightforward and does not require strong technical skills. It is possible to submit an annotation file either using the web browser or programmatically. Note that for programmatic upload Cloud Storage System drivers are available in different languages.
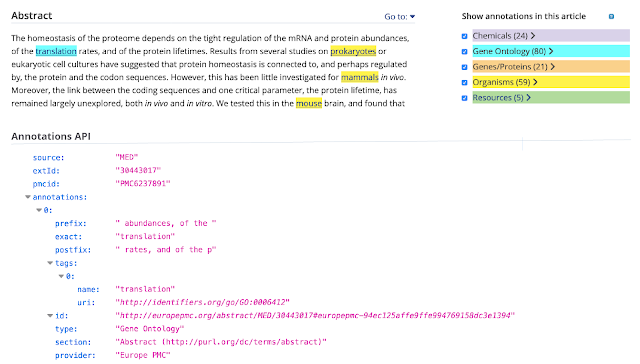
Here is how annotation data is represented in the system:
Europe PMC Annotation Submission System accepts sentence-based annotations and named entity annotations. An example of a sentence-based annotation is a protein-protein interaction, while a chemical name can be represented as a named entity. All annotated concepts must be linked to ontologies and data resources. For instance gene/protein annotations link to a corresponding UniProt record. Submitters are also asked to specify the precise location of the annotation in the text, using prefix and postfix tags.
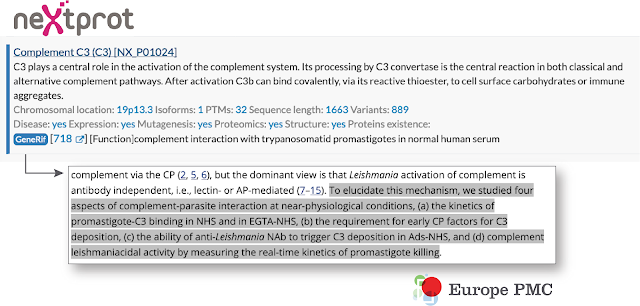
Such location information can be used to reciprocally link from the annotation to the relevant sentence of the article in Europe PMC via a link-back mechanism. neXtprot, an online knowledge platform on human proteins, participates in such link exchange. neXtprot entries often have associated functional gene annotations, known as geneRIFs. neXtprot users can now navigate from a neXtprot record directly to the relevant gene function statement found in the literature. Such location information can be used to reciprocally link from the annotation to the relevant sentence of the article in Europe PMC via a link-back mechanism. neXtprot, an online knowledge platform on human proteins, participates in such link exchange. neXtprot entries often have associated functional gene annotations, known as geneRIFs. neXtprot users can now navigate from a neXtprot record directly to the relevant gene function statement found in the literature.
Our aim is to make text-mining advances widely available for the benefit of the research community and we would not be able to do it without the support of our collaborators and annotation providers. Several text-mining groups have already made their annotations public in Europe PMC using the new submission system. Interested parties can share their results via Europe PMC platform, given that they adhere to the ground rules. If you would like to submit annotations please get in touch via annotations@europepmc.org. For more details join our free Annotation Submission Webinar on July 9th, 2PM GMT.