
The practice of preprinting in the life sciences has grown rapidly. In addition to accelerating scientific publication, preprinting also has the potential to open new avenues of communication among researchers. For example, preprint peer review offers tremendous potential for changing the culture of scientific assessment, broadening participation, and enhancing the robustness of scholarship. While only < 2% of preprints (themselves only posted for ~10% of biomedical papers) have accompanying reviews, this form of peer review is in the spotlight for many organizations engaged in advancing open science.
In order to support the growth of this emerging ecosystem, we must ensure that important metadata about preprint reviews are captured and shared with preprint servers and bibliographic databases in a machine-readable form. However, there is currently significant heterogeneity in the substance, format, and distribution channels of preprint review metadata.
At the ‘Supporting interoperability of preprint peer review metadata’ workshop, held on October 17 & 18 at Hinxton Hall, UK, and co-organized by Europe PMC and ASAPbio, representatives from preprint review projects, infrastructure providers, publishers, funders, and other stakeholders convened to collaboratively determine the key elements of preprint review metadata and mechanisms for sharing this information. Over two days, participants engaged in discussions on their visions of an ideal system, the metadata elements that should be included, the technical protocols and the next steps to take.
Day 1: Blue sky vision and prioritizing metadata
Blue sky vision
We kicked off the workshop with a series of four talks presenting blue sky visions of the ideal metadata ecosystem. This included contributions from Ludo Waltman (Leiden University), Martin Klein (COAR Notify), Thomas Guillemaud and Denis Bourguet (Peer Community In), and Michael Parkin (Europe PMC).
Ludo’s vision was to turn preprint peer review into truly meaningful bidirectional conversations and move away from it being simply a bureaucratic exercise. Martin’s vision from COAR’s standpoint was to develop an approach that is standards-based, interoperable, and decentralized to connect resources in a distributed network of repositories and external services. Peer Community In represented by Thomas and Denis shared their vision of communities of researchers handling the evaluation via peer review and recommendations of preprints in their scientific field, where readers are efficiently informed about preprint peer reviews and endorsements with the informed consent of authors. Europe PMC’s vision, represented by Michael, included a request for consistent use of persistent identifiers, clear licensing information to inform reuse, and standardized vocabulary of review types, to distinguish complete reviews from short comments.
Following that, in a breakout group participants discussed what they wanted to achieve by sharing preprint peer review metadata.
Some goals were shared by everyone:
Other goals spurred a lively discussion about associated nuances:

Prioritizing metadata elements for preprint peer review
The latter half of the first day focused on prioritizing metadata for preprint peer review. Participants examined 13 metadata elements identified as most important based on a questionnaire shared with a non-exhaustive list of metadata consumers prior to the workshop. These elements included:
- DOI of preprint reviewed
- Version of preprint reviewed
- Review date
- Reviewer name (or indication that review is anonymous)
- Recommendation/score
- Who selects reviewers (eg self-nominated or an editor/coordinator)
- Decision (binary or scalar)
- Reviewer competing interests
- Review license
- Review group name (eg for journal club or community)
- Inclusion of author response
- Reviewer ORCID
- Editor/coordinator competing interests
This session began with defining different use cases for these metadata elements and the benefits and difficulties of collecting them. Following in-depth discussions, individual participants ranked the 13 elements from the highest to lowest priority based on the perspective of the organization they were representing.
Metadata elements ranked with the highest to lowest priority from top to bottom:
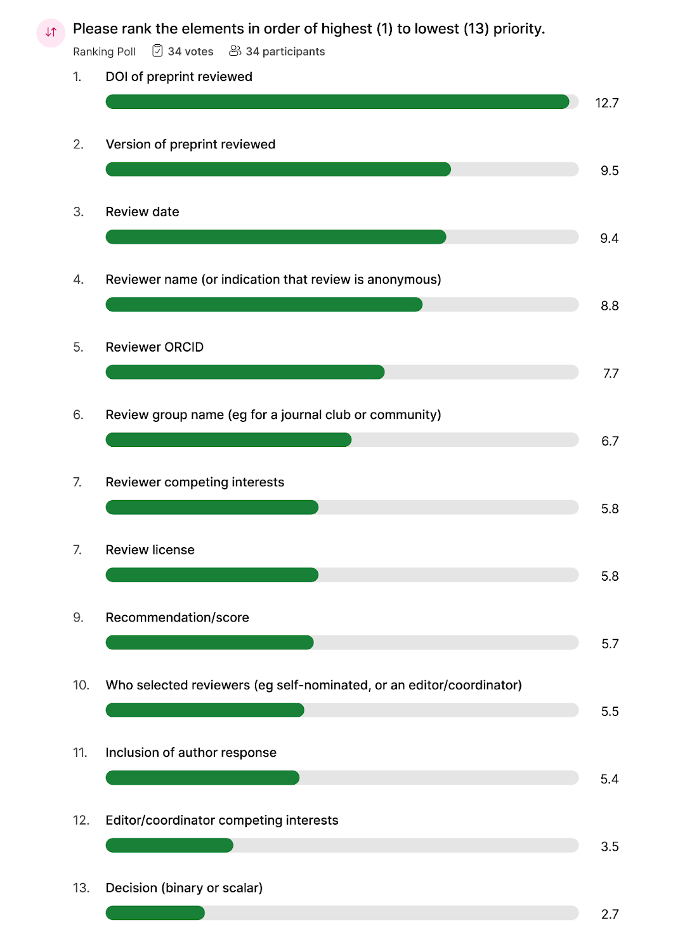
The DOI (or other PID) of the reviewed preprint was identified as the top priority, while the review decision (whether binary or scalar) was placed at the bottom of the list. The ranking exercise led to an animated debate that touched on several topics. Participants conferred if the reviewed preprint version must be encoded separately from the preprint identifier, depending on the versioning strategy of the preprint server. There was also a conversation around metadata describing reviewer expertise, a vague term that, despite potential for misuse, may nevertheless be desirable to some users. Capturing expertise could be particularly useful where reviewers wish to stay anonymous; otherwise reviewer’s ORCID record can be used to infer this information. Further key suggestions identified additional high priority metadata elements, including title of the review, review language for multilingual content, and ways to capture the complexity of different review groups and communities. Moving forward, we discussed transferring the ranking into core and desired to clearly identify metadata element requirements.
Day 2: Metadata transfer, scores & recommendations, and roadmapping
Metadata transfer pathways
The second day of the workshop began by focussing on the technical elements of metadata transfer pathways. This discussion examined the metadata elements that the various transfer protocols and services (including Crossref, DocMaps, COAR Notify, Sciety, DataCite, Zenodo, OA Switchboard and OAI PMH) currently share. Participants collectively discussed the barriers and benefits of different approaches before identifying significant gaps that exist for sharing preprint review metadata. In proposing solutions to these gaps, participants broadly agreed that more standardization and community agreement around a common approach would be beneficial.
Recommendation and scoring approaches
One of the most lively discussions of the second day was based around the inclusion and use of recommendations and review scores in preprint peer review metadata. The session began with an overview of some of the scoring methods (collated into this table), revealing a variety of different approaches that range from numerical to descriptive scoring. Participants created user stories for preprint review scores and decisions, which sparked a conversation around how this information would be used. While many participants thought that this metadata would be very useful, others were concerned that inclusion of scores in metadata could be manipulated by bad actors. Concerns primarily centered on the separation of a reductionist score from the larger review report. Many felt that even if a recommendation or score was beneficial, it should remain in the context of the larger review and reflected that by excluding scores from the list of minimal necessary metadata accompanying different user stories.
Roadmapping
The latter half of day 2 focussed on roadmapping activities to define concrete actions arising from this workshop. Participants began by individually listing next steps and activities prior to a larger conversation around the list. As part of this exercise, the list of potential actions was annotated according to importance and achievability. Following a second round of discussions on these prioritized actions, participants signed up to participate in on-going activities. Collectively, it was decided to create 3 working groups, each centered on a clearly defined action.
One working group is tasked with mapping the metadata transfer ecosystem. This group will map how services, depots and protocols interact to provide more clarity on how information can be shared.
A second working group will focus on developing a shared data model. This group aims to provide a common approach and protocols in addition to recommended standards, including defining the metadata elements. Additionally, this group will work to prepare metadata for sharing by different platforms, so as to reduce barriers to a common standard.
The third working group will further the discussion on recommendations and endorsements. This group will provide clarity on distinguishing reviews, endorsements and curation efforts. The group will also agree on recommended nomenclature and discuss the utility and risks of including recommendations within preprint review metadata.
Join us
Attendance at our workshop skewed toward organizations in the US and Europe, but development of the scholarly communication ecosystem should happen globally. As such, we are interested in broadening the conversation to include comments, suggestions, and participation from those with an interest in preprint review metadata; our working groups will periodically share project updates via a mailing list. If you would like to be part of this conversation please join via this form: https://forms.gle/g3FATYf57ASdQTUB9.
Co-authored by Summer Rosonovski and Johnny Coates



